Enzyme LSD1 found to regulate muscle fiber type differentiation
Our bodies convert food into energy for all life activities. These metabolic processes allow the cells that make up our bodies to produce (anabolism) and consume (catabolism) energy molecules from nutrients. In a recent update to a continuing research project from Kumamoto University, Japan researchers found that the LSD1 enzyme suppresses the expression of genes involved in mitochondrial metabolism and the slow-muscle genes that cooperate with them.All cells well regulate metabolism-related gene expression and adapt to environmental changes such as nutrition, oxygen supply, exercise, and temperature. Normal cells produce energy using mitochondrial metabolism when oxygen is abundant, and glycolysis (metabolism using mainly sugar) when oxygen is scarce. When metabolism changes between these mechanisms, the function of the cellular metabolic gene groups changes greatly. However, the mechanisms of these changes are still unknown.
Generally, gene function is determined by the state of transcription factors and genome modifications (chemical changes). The modified genome is called an "epigenome", and includes DNA methylation and modification of gene-regulating proteins, called histones, which DNA wraps around. Among these modifications, the methylation of histone lysine is an important part of the epigenome. It is created by methylation enzymes attaching methyl groups to histones and demethylating enzymes removing them.
In 2012, the Kumamoto University research group became the first to show the mechanism by which the demethylating enzyme LSD1 regulates energy metabolism in adipocytes (Nature Communications). Their research demonstrated the mechanism by which LSD1 suppresses the activity of mitochondrial metabolism genes, resulting in increases fat accumulation and obesity. Furthermore, in 2015, they revealed that LSD1 suppresses mitochondrial metabolism in cancer cells and promotes glycolysis in the presence of oxygen (Cancer Research). Thus, they showed that LSD1 is an epigenome modification enzyme that regulates metabolism depending on the state of various cells.
Skeletal muscle acts as a metabolic organ in the body and performs glycolysis in fast muscles and mitochondrial metabolism in slow muscles. In recent work, the Kumamoto University researchers used mouse myoblasts (C2C12 cells) to comprehensively analyze gene expression and the epigenome. They found that LSD1 suppresses both the expression of the genes involved in mitochondrial metabolism and the slow-muscle genes that cooperate with them. LSD1 was found to accumulate in metabolism-related loci and slow-muscle type loci to suppress gene function. Interestingly, the action of glucocorticoid, a major metabolic control hormone, promoted the degradation of LSD1 and reduced the LSD1 protein. In other words, myoblasts differentiate into fast muscle fibers in the presence of LSD1, and slow muscle fibers when LSD1 is suppressed with glucocorticoid. Thus, the relationship between the hormone glucocorticoid and the enzyme LSD1 was shown for the first time.
Researchers also found that a combination of glucocorticoid and LSD1 enhances the function of metabolism-related and slow-muscle type genes. Consequently, LSD1 coordinately controls skeletal muscle differentiation and metabolism under the action of hormones responding to the environment. This is thought to play an important role in skeletal muscle metabolism.
Recently, it has become clear that obesity and diabetes have common metabolic pathologies. Dementia, muscle diseases, and other age-related diseases also result from a functional decline of energy metabolism. In an aging society, the reduction of skeletal muscle mass (sarcopenia) and of mitochondrial metabolism in the skeletal muscle of obese and diabetic patients are drawing attention. The results of this research can be expected to be useful in the development of molecular mechanisms for new control and prevention methods of skeletal muscle dysfunction due to aging.
This research result was posted online in the journal "Nucleic Acids Research" on 29th March, 2018.
[Source]
Anan, K., Hino, S., Shimizu, N., Sakamoto, A., Nagaoka, K., Takase, R., … Nakao, M. (2018). LSD1 mediates metabolic reprogramming by glucocorticoids during myogenic differentiation. Nucleic Acids Research. doi:10.1093/nar/gky234
[Paper Info]
TITLE:
LSD1 mediates metabolic reprogramming by glucocorticoids during myogenic differentiation
AUTHORS:
Kotaro Anan, Shinjiro Hino*, Noriaki Shimizu, Akihisa Sakamoto, Katsuya Nagaoka, Ryuta Takase, Kensaku Kohrogi, Hirotaka Araki, Yuko Hino, Shingo Usuki, Shinya Oki, Hirotoshi Tanaka, Kimitoshi Nakamura, Fumio Endo, and Mitsuyoshi Nakao*
JOURNAL:
Nucleic Acids Research
(Oxford University Press)
DOI:
10.1093/nar/gky234
URL:
https://academic.oup.com/nar/advance-article/doi/10.1093/nar/gky234/4956186
[Fund]
(1) Japan Society for the Promotion of Science: KAKENHI,
(2) Takeda Science Foundation,
(3) Kanae Foundation for the Promotion of Medical Science,
(4) Ono Medical Research Foundation,
(5) Mochida Memorial Foundation for Medical and Pharmaceutical Research,
(6) Funding for open access charge: Takeda Science Foundation
<Fund numbers>
(1) JP15H04707, JP15K15068, JP16K07215, 25430178, (2) NA, (3) NA, (4) NA, (5) NA, (6) NA
[Image1]
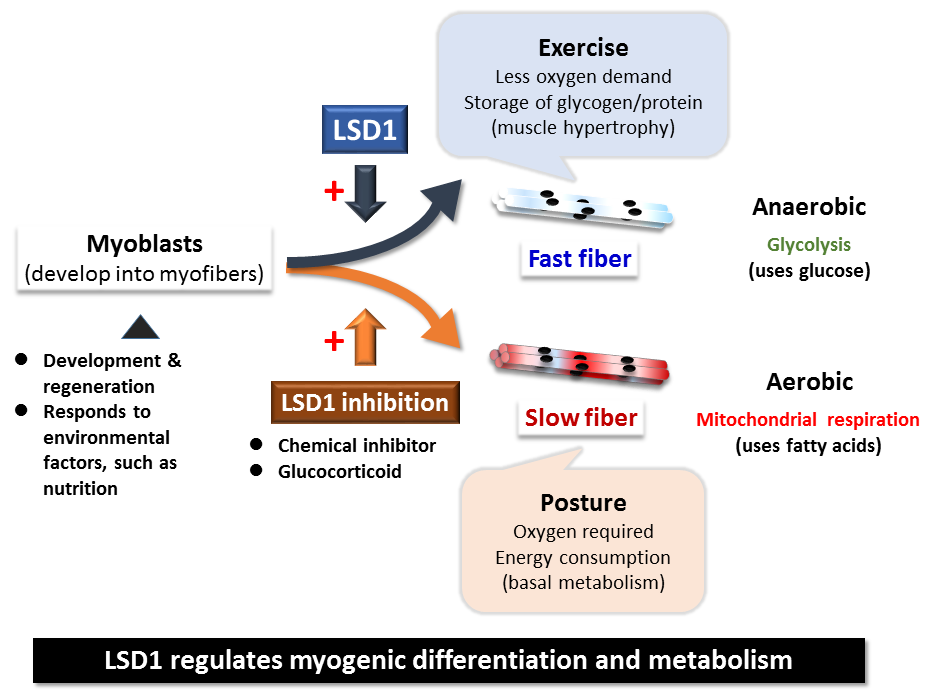
Myoblast regulation by LSD1
CAPTION:
The LSD1 enzyme regulates the differentiation and metabolic activity of skeletal muscle.
[Image2]
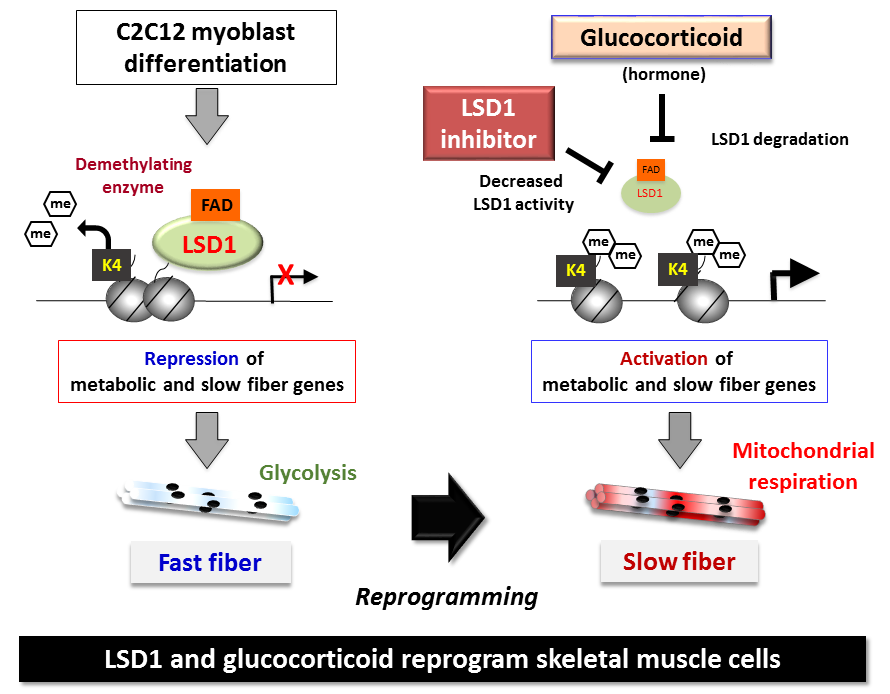
Skeletal muscle reprogramming with enzymes and hormones
CAPTION:
C2C12 myoblasts normally differentiate into fast muscle fibers in the presence of LSD1. Through the inhibition and degradation actions of LSD1 by a glucocorticoid hormone, the cells can be reprogrammed into slow muscle fibers.
[Image3]
Myoblast differentiation
CAPTION
Image of myoblasts differentiating into myofibers. Myoblasts can differentiate into two different types, slow-twitch (type 1) or fast-twitch (type 2) muscle fibers, each with distinct metabolic properties.
[Details for all images]
CREDIT: Prof. Mitsuyoshi Nakao
USAGE RESTRICTIONS: These images may only be used in conjunction with the accompanying release, or stories written about the work described in the release with reference to the original work.
